NMsim is an R package that can simulate Nonmem models (using the NMsim function) based on just a simulation data set and a path to an estimation control stream. It will also retrive and combine output tables with input data once Nonmem has finished and return the results to R.
NMsim is on CRAN, MPN and github:
## From CRAN/MPN repositories
install.packages("NMsim")
## From github
library(remotes)
install_github("philipdelff/NMsim")In its simplest use, a simulation of the (estimated) model stored in “path/to/file.mod” using the simulation input data set stored in the variable data.sim this way:
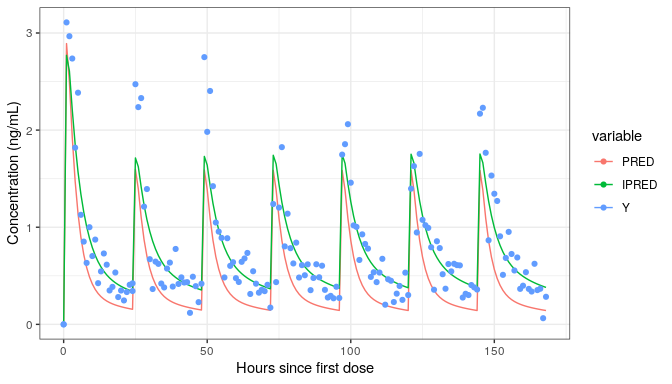
We are ready to plot
library(ggplot2)
datl <- as.data.table(simres) |>
melt(measure.vars=cc(PRED,IPRED,Y))
ggplot(datl,aes(TIME,value,colour=variable))+
geom_line(data=function(x)x[variable!="Y"])+
geom_point(data=function(x)x[variable=="Y"])+
labs(x="Hours since first dose",y="Concentration (ng/mL)")
This example was a simulation of a multiple dose regimen with a loading dose using a model estimated on single dose data. It is from the first vignette NMsim-basics.html.
NMsim has a flexible way to define simulation methods. The following methods are currently provided:
method.sim=NMsim_default)method.sim=NMsim_typical)method.sim=NMsim_known)method.sim=NMsim_VarCov)method.sim=NMsim_asis)To learn how to run these simulations on your Nonmem models, get started with NMsim-basics.html. It is really easy.
In addition, NMsim can simulate multiple models at a time. E.g., if a bootstrap run of a model is available, NMsim can run the simulation with each of the bootstrap models and collect all the results in one dataset. This provides a robust and easy way to simulate a Nonmem model with uncertainty.
You can also write your own methods, if you have some other Nonmem-based simulation (or other job) you want to automate using NMsim.
Many features are available. Prominent ones are:
SUBPROBLEMS feature avaible through the subproblems argumenttransform argument.If residual variability is not implemented in the simulated model, NMsim provides a way (addResVar()) to add residual variability in R after the simulation has been run.
One strength of NMsim is that it does not simulate, translate or otherwise interpret a Nonmem model. Instead, it automates the Nonmem simulation workflow (including execution of Nonmem) and wraps it all into one R function. In the example given above, NMsim will do the following:
file.mod ($INPUT and $DATA matching the saved simulation data set; $SIMULATE instead of $ESTIMATION and $COVARIANCE)file.ext)This eliminates the need for re-implementation of a model for simulation purposes. On the other hand, this also means that NMsim can’t work without Nonmem.
NMsim can call Nonmem directly or via PSN. If NMsim is run on a system where Nonmem cannot be executed, NMsim can still prepare the simulation control stream and datafile.
NMsim is in itself a small R package. It makes extensive use of functionality to handle Nonmem data and control streams provided by the R package NMdata.
The methods currently provided by NMsim will work with (many or most) Pop PK models and most continuous-scale PD models. Methods are currently not provided for for time-to-event models. Also, depending on the coding of the models, other censored data models may not work out of the box, because the model may not have a single variable (in Nonmem) that simulates the wanted information for all data rows, as their interpretation may depend on other values.
The input data set must contain whatever variables are needed by the Nonmem model. A common issue is if the Nonmem model uses a covariate that is not in the simulation input data set. NMdata’s NMcheckData is a good help identifying input data issues before running Nonmem - and when Nonmem acts unexpectedly.
Nonmem may not be the fastest simulator out there. But actually most often, the reason Nonmem is slow at providing a simulation result is that it takes a long time writing the $TABLE files (yes, that can account for 90% or more of the time Nonmem spends). NMsim provides a simple way to get around this. The argument text.table can be used to define only the columns needed in the simulation output (which may be as little as PRED, IPRED, and a couple more - remember the input data is merged back automatically). As a result, NMsim may still be slower than a re-implementation in a different framework. But it’s extremely easy to do.
Currently, NMsim can only run Nonmem on Unix/Linux systems. It wouldn’t be too big a deal to add support for Windows, so reach out if you need it. It is possible to run R on Windows and run Nonmem on a Unix/Linux system through SSH if needed.
NMsim does not need PSN but can use it. However, not all features are available with PSN, so for some features you will have to specify the path to the Nonmem executable (say path.nonmem=/path/to/nmfe75 or any Nonmem executable you want to use). Specifically of the simulation types currently available, simulation of known subjects is not possible using PSN (but works if a Nonmem executable is provided).
If PSN is used, NMsim uses PSN’s execute to run models. In addition, NMsim by default uses PSN’s update_inits to update initial values in control streams, if PSN is available. NMsim does also include its own simple function to do this if PSN is not available.
NMsim reliable?Importantly, NMsim does not (at least not by default) modify, translate or simulate the model itself. It does modify control stream sections $INPUT, $DATA, $ESTIMATION, $SIMULATION, $THETA, $OMEGA, $SIGMA, $TABLE as needed. The fact that NMsim allows for skipping the re-implementation but just uses Nonmem to simulate the Nonmem model as is, eliminates the risk of discrepancies between the estimated model and the simulated model.
The produced control stream is saved together with simulation data set open for manual inspection and can obviously be run with Nonmem independently of NMsim.
NMsim includes functions (NMcreateDoses and addEVID2) to very easily create simulation data sets. While one certainly does not need to use these functions to use NMsim, they do add to the package providing a framework that enables a complete simulation workflow in only 5-15 simple lines of R code.
There are several other packages out there that can do this, and NMsim may not be your best choice if this feature is all you are looking for. However, running Nonmem using the NMexec() function provided by NMsim has one important advantage in that it saves the input data together with the Nonmem control streams. This ensures that output data can be merged with input data as it went into the model, even if the input data file should be modified or lost.
NMexec will submit model runs to a cluster by default. This can be switched off for running Nonmem locally. Please notice the jobs are submitted to a cluster in a very specific way using PSN. If your setup is different, this is for now not supported. Please use NMexec(sge=FALSE) in that case (which may not be desirable). Notice that simulations are not done on a cluster by default so you may still be able to use NMsim.